About Me
I'm a Bioinformatics Engineer currently working at Agroscope (Switzerland) on de novo genome assembly and genetic diversity in grass ecotypes. I recently completed my M.Sc. in Bioinformatics from the University of Rouen, with an 18-month apprenticeship at INRAE.
My work centers on pangenomics, genome assembly, and NGS analysis, where I develop computational tools and workflows for large-scale genomic data. These are topics I'm eager to explore further and at greater depth.
I also served as House Manager at the Bordeaux National Opera for five years, coordinating reception teams during evening performances, a role that taught me rigor, teamwork, and attention to detail.
Experience
Bioinformatics Engineer Intern
Agroscope - Wädenswil, Switzerland
De novo genome assembly and genetic diversity analyses in Swiss ecotypes of Lolium multiflorum.
Bioinformatics M.Sc. Engineer Apprentice
INRAE BIOGECO - Cestas, France
Developing pangenomics tools and workflows to help identify structural variants in Quercus spp..
Proteomics B.Sc. Research Intern
INRAE BFP - Villenave-d'Ornon, France
Statistical analysis of fruit proteome across 5 species, identifying differential expression patterns across developmental stages.
Education
M.Sc. Bioinformatics, Modeling and Statistics
University of Rouen Normandy
2nd year of M.Sc. 18-month apprenticeship program
Deep Learning Research Exchange
University of Groningen - Netherlands
Deep learning of satellite imagery applied to urban forestry
M.Sc. Bioinformatics
University of Bordeaux
1st year of M.Sc.
B.Sc. Life Sciences
University of Bordeaux
Main skills
Bioinformatics
- Genome Assembly ⋆
- Pangenomics ⋆
- NGS
- Workflow Development
- Structural Variants
Programming
- Python ⋆
- Snakemake ⋆
- Bash
- R
- LaTeX
DevOps & HPC
- Apptainer ⋆
- Cluster Computing
- SLURM, Hetzner
- Git
- CI/CD
Projects
MSpangepop ⋆
Python library for pangenome simulation under coalescent demographic models. Features clean, object-oriented architecture designed for extensibility and ease of use by biologists.
Supervised by : DSc Ludovic Duvaux

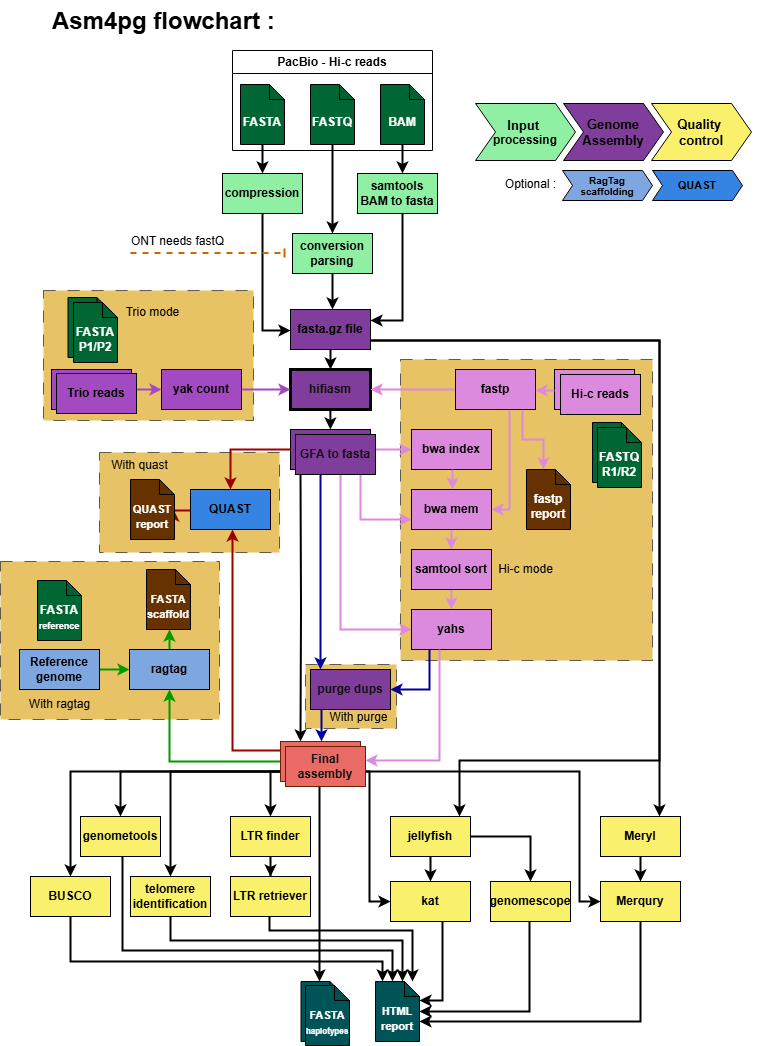
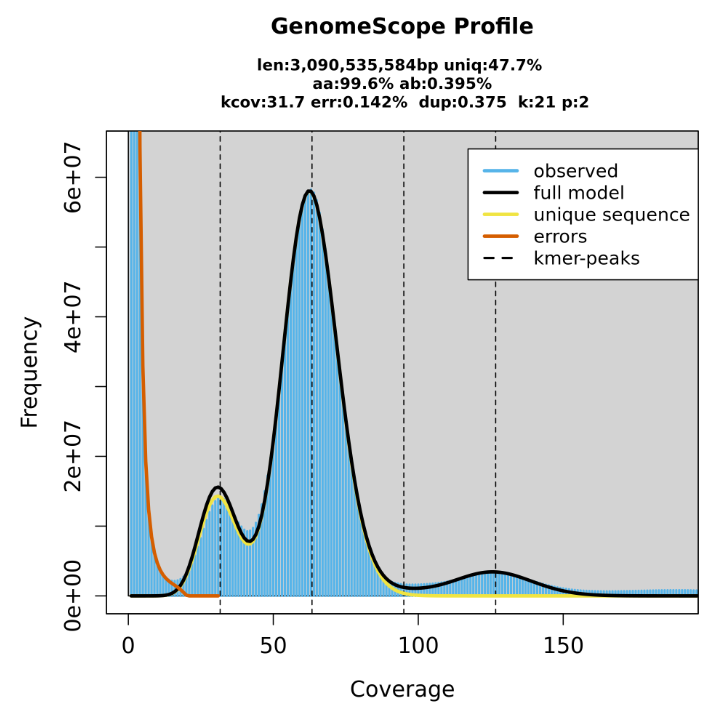
Asm4gp ⋆
Comprehensive long-read assembly and QC pipeline supporting PacBio, ONT, TRIO, Hi-C, and HiFi data. Developed using Snakemake to automate haplotype-resolved genome assembly inputs for pangenome construction.
Supervised by : DSc Ludovic Duvaux
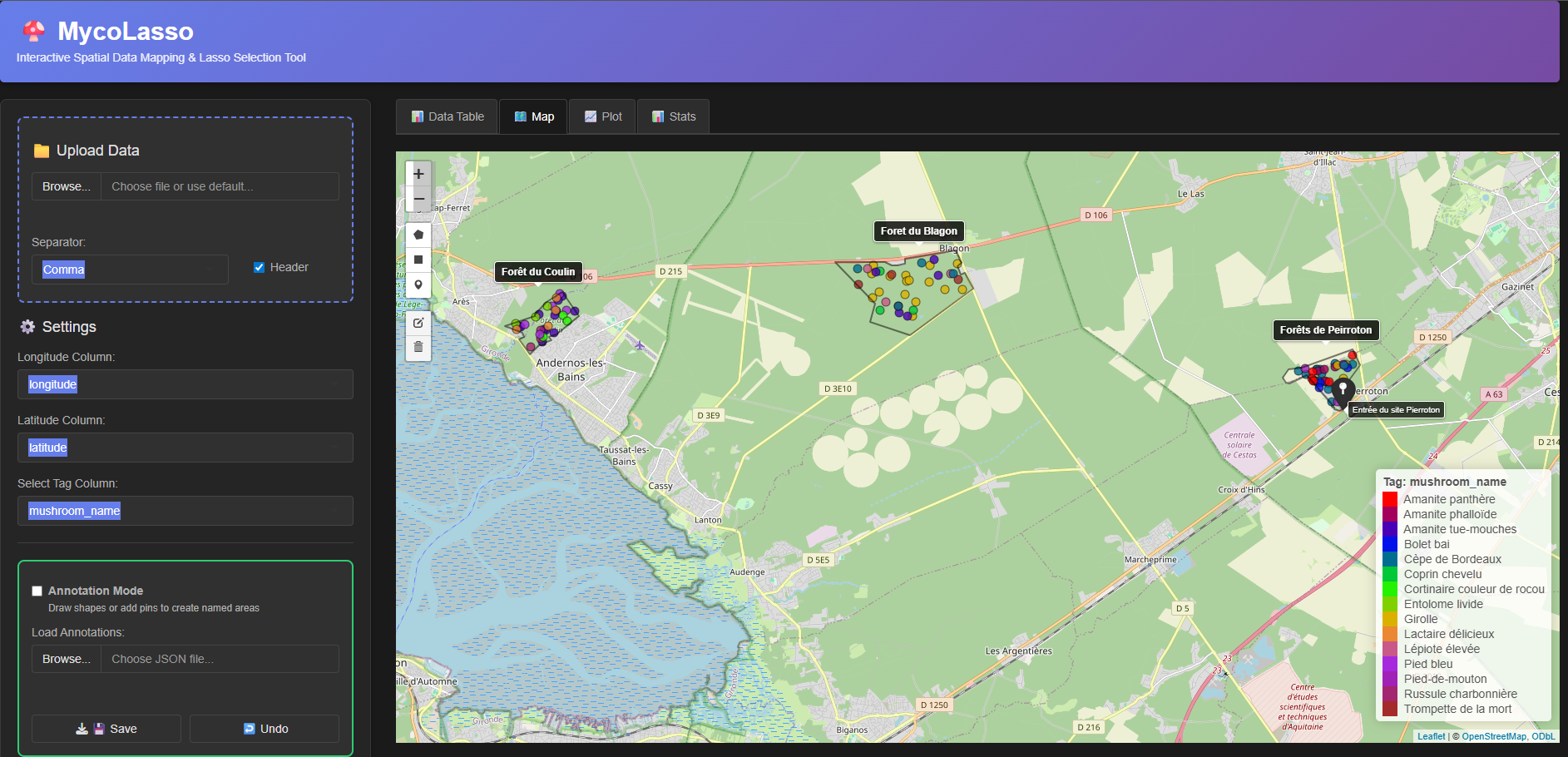
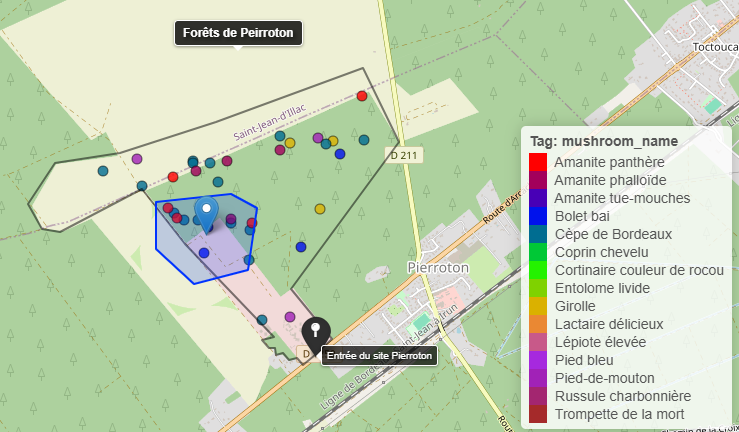
MycoLasso
MycoLasso is a Shiny app to load, visualize, and interactively explore geospatial data points.
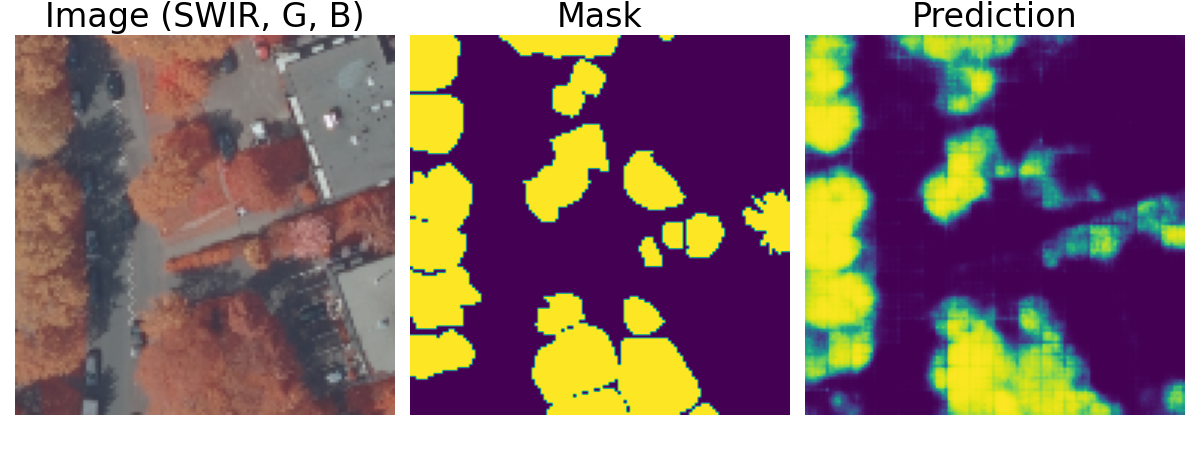
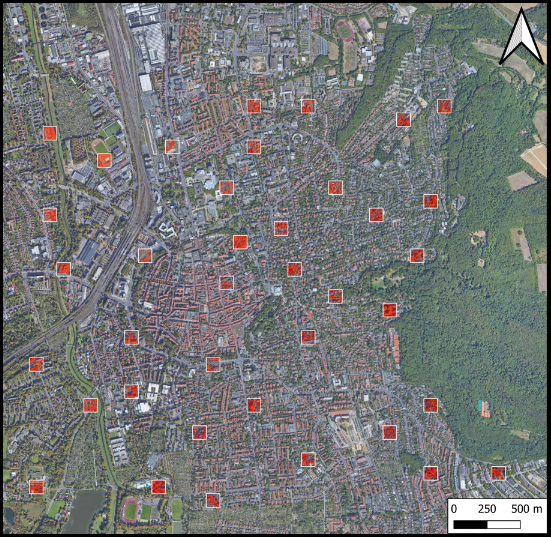
Urban Tree Detection
Deep learning pipeline for automated tree detection and counting from satellite imagery using U-Net architecture. Developed during Erasmus+ exchange at University of Groningen.
Supervised by : Pr Jean-Christophe Taveau

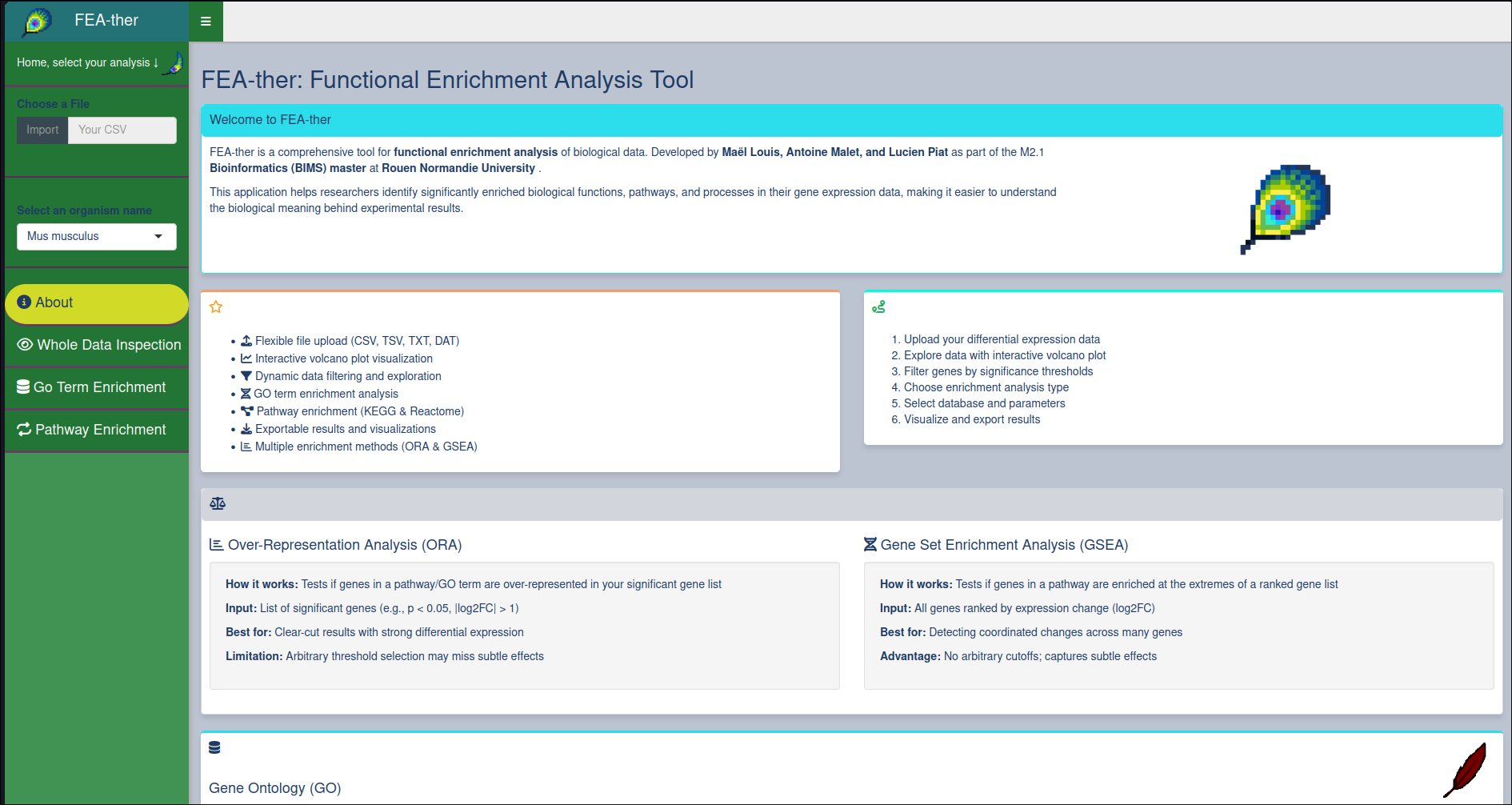
FEAther
Shiny application designed for functional enrichment analysis of biological data. It allows users to upload CSV files, select various analysis options, and visualize results through interactive plots and tables.
Supervised by : Asst. Prof Hélène Dauchel

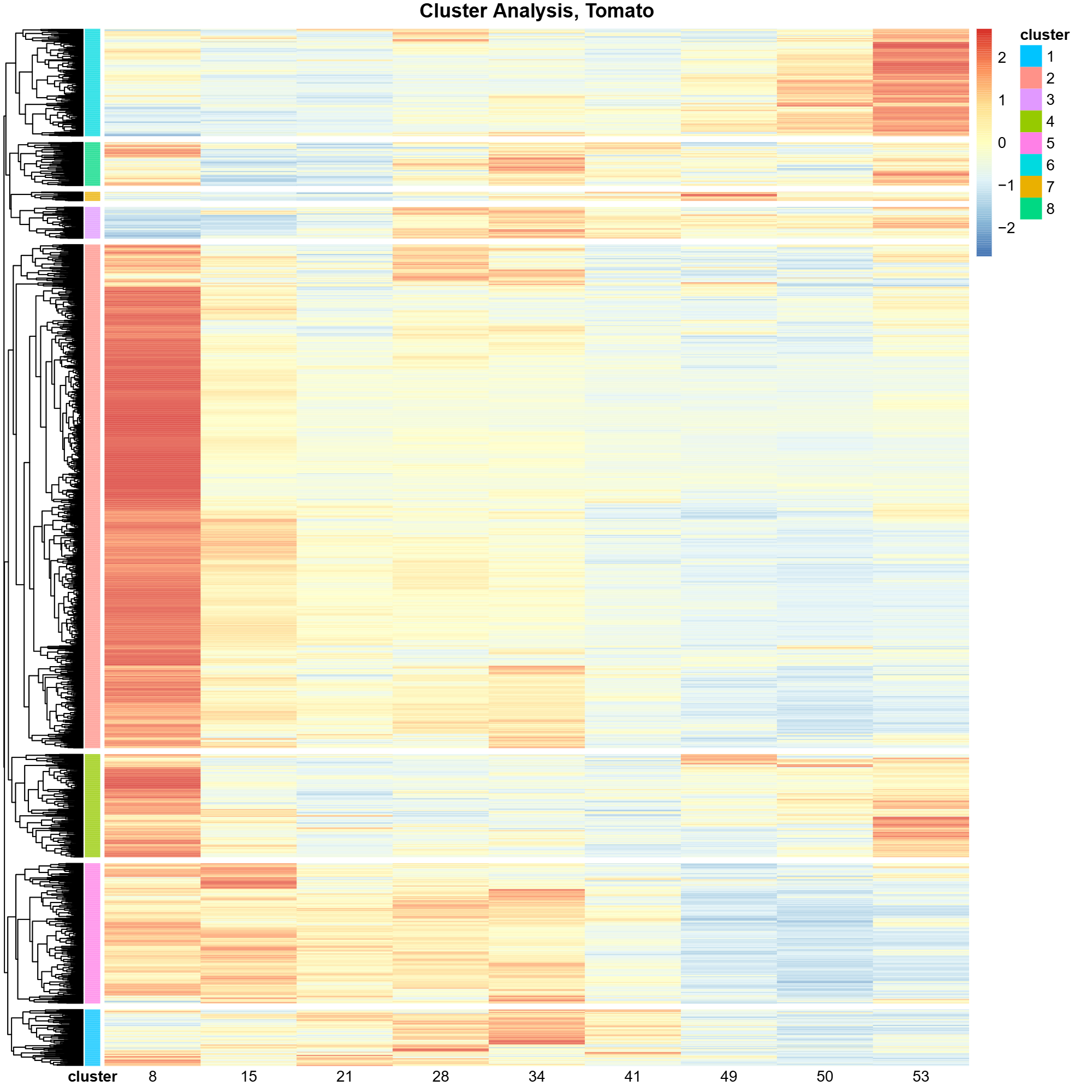
Fruit Proteomics Analysis
Statistical analysis of proteomic data across five fruit species to identify differential protein expression patterns during development. Utilized R for data processing, visualization, and interpretation.
Supervised by : Dr Sophie Colombié

Publications & Reports
Papers
- MSpangepop: Simulating complex structural variants under advanced demographic scenarios using the coalescent JOBIM 2025, Bordeaux, France HAL
- The pangenome of European white oaks: A new approach to identify potential barriers to gene flow EVOLTREE & Forgenius Final Conference, Nov 2025, Madrid, Spain HAL
- The pangenome of European white oaks: A new approach to assess genetic diversity Doctorales de la foret, Feb 2025, Poitiers, France HAL
Reports
- De la construction a la simulation des pangenomes : une approche bioinformatique integree via Asm4pg et MSpangepop PDF M.Sc. apprenticeship report, INRAE BIOGECO / Univ. Rouen, 2024-2026 | Supervised by Dr. Ludovic Duvaux
- Single Tree Detection: The Jurassic Bark PDF Erasmus+ research project, University of Groningen, March 2024
Contact
Feel free to reach out for collaborations or opportunities!
lucienpiat33@gmail.com